by Jared Smith
Please contact me if you have any corrections or clarifications.
This document WILL:
This document WILL NOT:
This guide assumes you have uploaded your DNA data to GEDmatch.com and that it has been fully batch processed. This means your DNA data is in the GEDmatch database so it can be used to compare to others. If you haven't uploaded yet, please follow the instructions at GEDmatch to upload your file(s), then wait a day or two until the analysis has completed.

Chromosomes are tiny structures found within your cells. They contain the DNA information and instructions that define who you are - what you look like, how your body works, and even what genetic diseases you might have.
Humans have 46 chromosomes. But chromosomes come in pairs, so we typically think of them as 23 pairs of chromosomes. The first 22 chromosome pairs (called autosomes) are numbered 1 through 22. We'll primarily focus on these autosomal chromosomes. The 23rd pair are called the sex chromosomes - men have an X and a Y sex chromosome and women have two X chromosomes.
One autosomal chromosome from each pair comes from your mother and the other comes from your father. This means you get half of your DNA from your mother and half from your father. Each chromosome they pass on to you is a combination of their own pair of chromosomes which they got from their parents (your grandparents).
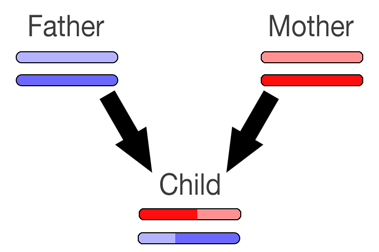
The image above depicts how one pair of chromosomes may be passed from your parents to you. The colors don't mean anything special - they simply depict the individual chromosomes and chromosome sections.
You'll notice that the chromosome passed to you from each parent may not be an exact 50/50 combination of their own chromosomes. This means that you might have a bigger portion of one of their chromosomes than the other - you might be more related to one of your grandparents than another on that chromosome. In fact, you might have an exact copy of one of your parent's chromosomes, and thus you'll get no portion of their other chromosome.

If you add in another generation, things get a bit more complex. This depicts just one chromosome pair. Remember that you have 22 pairs that will be various combinations of your grandparent's chromosome pairs. In this example, one of the mother's chromosomes (the one she got from her father) was passed on directly to the child. This child will not match his maternal grandmother on this chromosome. I'm not sure how often this non-recombination occurs, but of my 44 autosomes, 6 were not recombined from my parents to me. While lop-sided chromosomes or non-recombination may occur on a particular chromosome, across all 46 chromosomes, things tend to average out - you'll get around 25% of your DNA from each of your grandparents.
For each generation you go into the past, you will get less and less of that ancestor's DNA. The chromosome segments they pass on will become smaller or lost due to recombination. This is why autosomal DNA analysis is usually only useful to at most 6 or 7 generations back - you have so little DNA from very distant ancestors that it becomes difficult to analyze it reliably.
Centimorgan (abbreviated cM) is a measure of genetic linkage. Think of it as a measure of DNA information within a chromosome. Each chromosome contains different amounts of information. Chromosome 1 contains 281.5cM of information. Chromosome 2 has 263.7cM. Chromosome 21 has only 70.2cM.
SNPs, or single-nucleotide polymorphisms, are tiny pieces of a chromosome that contain distinct blocks of information. There are thousands of them per chromosome. SNPs are compared between two people to see if they match. The amount of information in matching SNPs is measured in cM.
The cM values for SNP matches are sometimes referred to as "chromosome length" or "match length". However, information is more densely packed in certain areas or SNPs within chromosomes, so there's not a direct correlation between number of SNPs and cM amount. When you view GEDmatch's graphical depiction of chromosome matches, a bigger matching block does not always mean a higher cM value.
A "segment" refers to a section or block of contiguous SNPs. A "matching segment" is a section that is the same between two people.
Individual markers (called base pairs - the things that SNPs are made of) within a chromosome are numbered. There are millions of these markers per chromosome. A segment of a chromosome can be identified by these location numbers.
Sometimes SNPs marker values match between two people simply by chance. This is called IBS or Identical By State. And sometimes they match because they were passed down from a common ancestor. This is called IBD or Identical By Descent.
This is Most Recent Common Ancestor - the ancestor from which you and a DNA match received your common DNA segments.
Using the terms above, you can begin to speak the language of genetic genealogy. For example, you may have a match with another person on a segment of Chromosome 3 from marker start location 36,495 to end location 5,168,135 for a total of 15.8cM of information in 2,114 matching SNPs.
GEDmatch can show these types of matches in a table and with a graphical representation of the chromosome:
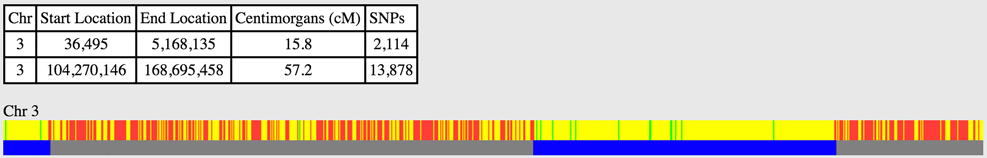
The blue bars indicate two segments that match on Chromosome 3 between two people. The table indicates Start and End Locations and the cM and number of matching SNPs in each segment. You'll notice that the start location for the first segment is 36,495 instead of 0 even though it appears at the beginning of the chromosome - this is because not all markers in a chromosome, especially those near the ends, are tested. We'll discuss the other colors in this graphic later.
The larger the segment (more SNPs and higher cM) of matching markers/base pairs, the more likely it is IBD (you share a common ancestor) rather than IBS (just matching by chance). Matching segments smaller than 7cM or 700 SNPs have a high likelihood of being IBS, so they should be considered questionable. Matches smaller than 3cM or 300SNPs should be highly suspect and rarely used alone for genetic genealogy.
If you add up the total of all cM values for the segments someone shares with you, you can get a rough calculation of how closely you are related to them. There is a total of around 6800cM in all 44 autosomal chromosomes. The following are expected cM matching values for various relationships:
The cM match amount or overlap decreases as your relationship gets more distant. You might share only 13cM (.195%) with your fourth cousin (someone with whom you share a 3rd great-grandparent). Of course with the variability of many generations of recombination (or non-recombination) of chromosomes, you could share much more than that, or you could share 0cM and not be identified as a cousin match at all.
You can get a full table of expected cM match values for various relationships at http://isogg.org/wiki/CentiMorgan.
There is much variability in DNA tests. Each company tests slightly different things in different ways. DNA inheritance is highly variable. For all of these reasons, keep in mind that the cM match values and predicted relationships are VERY ROUGH ESTIMATES ONLY!.
This is especially true for more distant cousins. Additionally, if you are related to someone on multiple lines - or if you or your match are related to your common ancestor on multiple lines (e.g., your grandparents were cousins) - then the total cM will suggest a closer relationship than is actually the case.
The One-to-many Matches report will provide a list of people you share chromosome segments with. To view the report, click the 'One-to-many' matches link on the home page and select your kit # (found on the homepage) on the next page. We'll be comparing Autosomal chromosomes, not X, so make sure Autosomal is selected. Keep threshold at 7 cM and select Display Results.
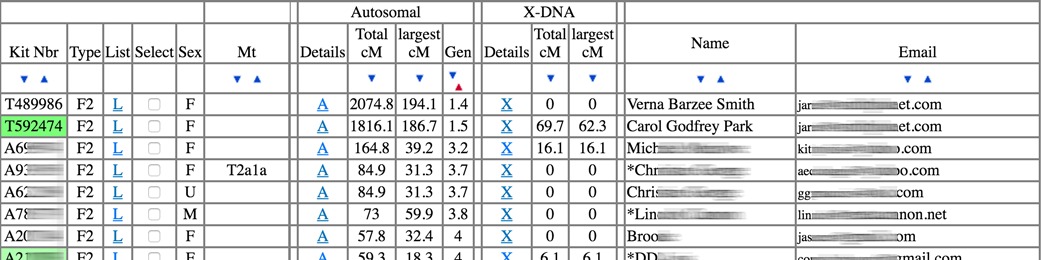
The large table will list your matches in order of Total cM overlap. Most everyone on the list (especially those near the top) will be related to you... somehow. The report also displays the largest cM segment amount you share. The Gen column provides a rough estimate of the number of generations between you and the Most Recent Common Ancestor (MRCA) you and that match both share - 1 for parent-child, 2 for 2 generations (grandparent-grandchild), etc.
In the screenshot above, the top two results are my grandmothers. Notice their Gen values are 1.4 and 1.5 - I got a bit more than 25% of my DNA from each of them. The next several results are all known cousins of mine - generally in the 3rd cousin range. Gen of around 3 suggests common great-grandparents (probably around 2nd cousins), Gen of 4 suggests common great-great-grandparents (around 3rd cousins), etc.
Kit Nbr provides an identifier for each person you match. The beginning letter indicates the testing system they used - T = Family Tree DNA, A = Ancestry.com, and M = 23andMe.
Clicking the "L" in the List column will run a One-to-many Matches report for that person. This can be handy to see who that person matches.
Clicking the "A" in the Details column will run a 'One-to-one' compare report between the person whose matches list you are viewing and the person listed in that row.
The other columns are either self-explanatory or are not relevant to this discussion. Names that begin with a * in the Name column are alias names and may not be the actual match's name.
You should regularly monitor your matches list for new DNA cousins. Newly added kits show with a green background. I recommend keeping a spreadsheet or document with information and notes about your cousins - especially ones with whom you have identified your relationship and common ancestor.
GEDmatch uses a batch process to generate your One-to-many matches list. It can sometimes display people you aren't actually related to. Be sure to do a One-to-one compare with a listed match to ensure you actually share matching DNA segments.
We will not explore X Chromosome analysis in depth here, but keep the following in mind:
You can read more about X Chromosome matching at http://smithplanet.com/stuff/x-chromosome.htm.
The One-to-one compare utility allows you to look for chromosome segment matches between two people. You can run this utility by selecting 'One-to-one' compare on the homepage and entering the kit #s for the people you want to compare, or by clicking the "A" link on the One-to-many report. The default settings will generally suffice for most matches, though I prefer to enable the Show graphic bar for each Chromosome? option to give a more visual presentation of the segment overlaps.
Remember, our chromosomes come in pairs. However, when the DNA testing tools do chromosome comparisons, they can't distinguish between the two pairs of a chromosome - they instead treat them essentially as one combined chromosome - as if the chromosomes have been laid on top of each other.  This means that when you match someone on a chromosome segment, you can't be sure which of your chromosomes they match. It could be the chromosome you got from your father or the one you got from your mother.
This means that when you match someone on a chromosome segment, you can't be sure which of your chromosomes they match. It could be the chromosome you got from your father or the one you got from your mother.
Think of this like looking through a double-pane window. When you see a streak on the window, it's difficult to tell whether it's on the inside pane or the outside pane without analyzing it very closely or from another angle. The same applies to DNA matches - you must analyze them closely or compare them with someone else in order to know what they mean.
If you match two different people on the same, large (7cM+) chromosome segment, those two people are related to you, but they may not share a common ancestor or be related to each other. You must always do a One-to-one compare between your matches to make sure they also match each other on the same segment(s).
A half match indicates that an SNP of one of your chromosome pairs matches the corresponding SNP in one (or the other) of someone else's chromosome pairs. Half matches are depicted in yellow in the chromosome graphic bar.
When the SPNs for both of your chromosome pairs are identical to both pairs from someone else, this is called a full match. Full matches display in green in the chromosome graphic bar. Full matches on large segments are not common - typically only in twins and full siblings, or when someone's parents are related (as is common in endogamous cultures or regions). The small green lines in the graphic below indicate SNPs that match on both chromosomes simply by chance (IBS).
The red lines below indicate there is not a match on these SNPs. The yellow lines interspersed with the red lines are IBS matches - the SNPs match, but only by chance. The blue bars indicate large segments (>7cM by default) that are half or full matches. Because DNA tests report only the total amount of DNA where either (half match) or both (full match) of the pairs are the same, identical twins will show as sharing 3400cM of DNA, rather than the actual 6800cM.
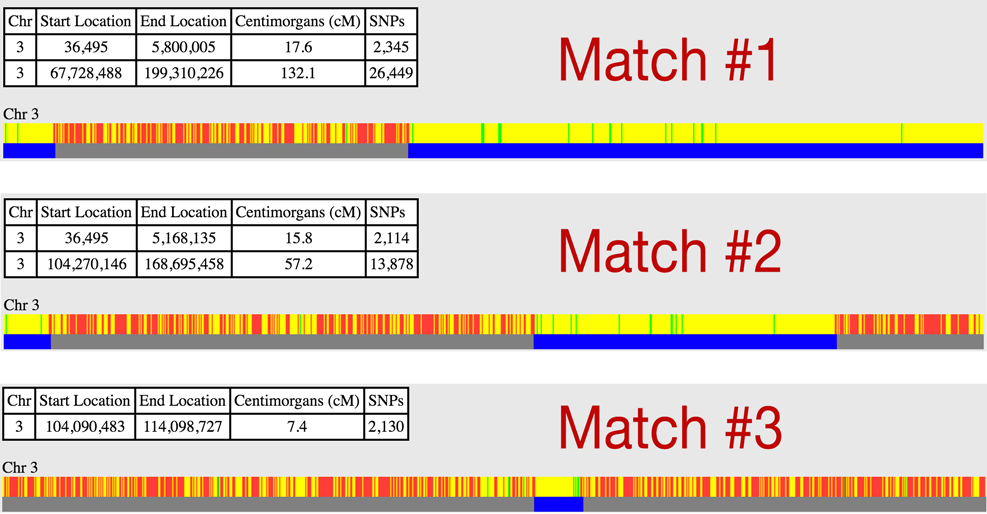

The graphic above shows the Chromosome 3 segment overlaps between me and 3 separate matches. At first glance, you might assume that all three of them are related - they each share notable overlaps in the same areas of my Chromosome 3. All four of us must have received the matching segments from a common ancestor, right? WRONG!
In this case, Match #1 is my paternal grandmother and Match #2 is my maternal grandmother. They do not share any DNA with each other and are not related (except for sharing me as a grandson)! They appear to be matches because we're actually comparing both of their Chromosome 3s to both of my Chromosome 3s. My paternal grandmother matches these sections of one of my chromosomes and my maternal grandmother matches these sections on my other chromosome. The fact they match in similar areas is simply coincidental. A One-to-one compare between my grandmothers proves there is no match and my grandmothers are not closely related (at least on this chromosome):

As before, the small half and full match lines are IBS only because these segments are so small.
Let's now analyze Match #3. The process of determining if a match is related to another match is called triangulation. We know all three matches are related to me, but we want to triangulate to see if Match #3 is related to one or both of my grandparents.
I only have two Chromosome 3s - and the area in which Match #3 matches me is the same area in which I match my grandmothers - so we can be assured that Match #3 is thus related to one of my grandmothers. But which one? We can determine this by doing a One-to-one compare between Match #3 and both of my grandmothers. Comparing Match #1 (my paternal grandmother) and Match #3 shows no matching segments. We therefore know that the match is with my maternal grandmother (Match #2)! This is what we see when we One-to-one compare them:

This is definitely a match! They match each other, and both of them match me on the same segement. In this case, Match #3 is my maternal grandmother's first cousin (my first cousin twice removed).
You can see that my grandmother and her cousin share much more of Chromosome 3 than I share with her cousin. This is to be expected - Match #3 is more closely related to my grandmother than to me. This also indicates that the portion of Chromosome #3 that they share, but that I don't share with my maternal grandmother was not passed on from my grandmother to me via my mother. So, if I match a cousin in the area of Chromosome #3 I didn't get from my grandmother, then I know I match them on either my maternal grandfather's line or one of my paternal lines. Eliminating lines for possible MRCAs can be very valuable.
You can use this type of logic and analysis to slowly build a mapping or spreadsheet of all 23 of your chromosome pairs. As you establish your relationship to cousins, you can begin to identify whether a match is on your mother's or father's side, and then which two of your four grandparents a particular chromosome pair segment maps to. If you have identified a common ancestor with a cousin for that segment, then matches on that chromosome which also triangulates as a match to another known descendant of that ancestor will also share that common ancestor (or perhaps an ancestor or descendant of that ancestor). If you don't have known cousins with which to triangulate, you have to be careful in making assumptions - the match could be on either of your chromosomes and on any of your family lines.
The more cousins you identify common ancestors with, the easier it becomes to identify common ancestors with additional cousins. Start at the top of your matches list and start contacting matches (be sure to One-to-one compare first!) and slowly build a list or spreadsheet of your cousins, the chromosome segments you share, and your common ancestors.
This very valuable GEDmatch tool allows you to more easily identify cousins who are related to each other. It is often called the "In Common With" (ICW) tool. This tool shows you those who are (and are not) related to two different people. If you have identified a cousin, run this tool on your kit # and their kit # to find people who are related to both of you.
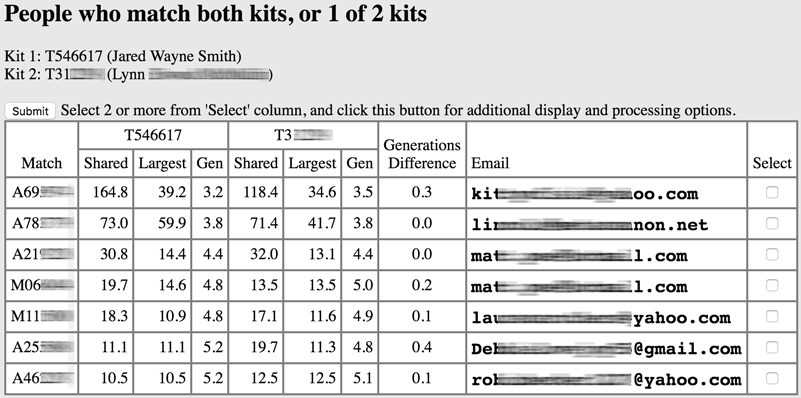
You can now analyze these common matches to verify (or refute) your relationship via triangulation. The Gen columns provide estimates of the distance to the MRCA for the two people you're comparing and the common match. Differences in these values may suggest that one of you is more closely related to the MRCA than the other (i.e., you are probably cousins once or twice removed). As before, GEDmatch can't differentiate between your chromosomes, so make sure you do One-to-one between everyone involved before assuming that matching segments mean you share a common ancestor.
Each child gets an entirely different combination of their parent's chromosomes (unless they are identical twins). This means that you can match with someone that a sibling (or other relative) does not.
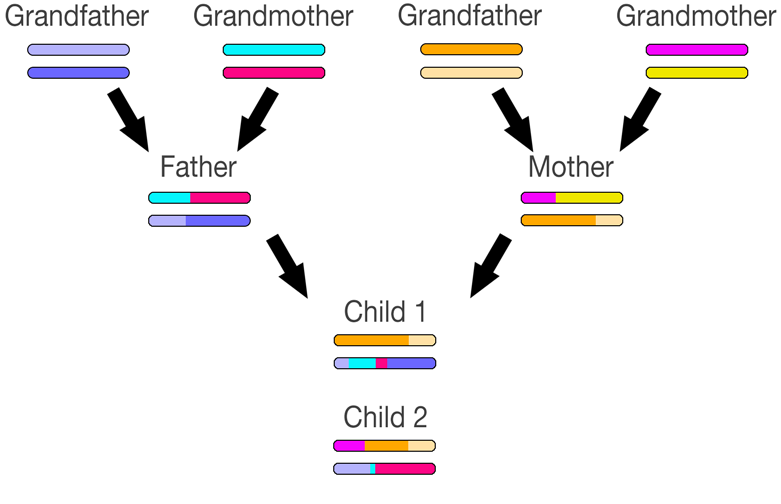
In this example, you can see how the children match different segments of their grandparent's chromosomes. A match with a distant cousin in the dark blue section for Child #1 would not be present at all for Child #2. And Child #1 wouldn't match any relatives on the maternal grandmother's line on this chromosome due to lack of recombination.
If you don't match someone, this does not always mean you are not related. It's possible that matching segments from an ancestor simply weren't passed down to each of you. A match to one of your known relatives may still be your cousin, even if you don't match them - especially if their segments align on your chromosome map to your ancestor. Testing additional relatives can be helpful in finding additional cousins and unknown ancestors. They will establish a family baseline with which to triangulate to determine genetic ancestry lines.
Testing older relatives (especially parents and grandparents) will get you an extra generation (or 2 or 3) further back in time - enough to discover cousin connections that would otherwise be impossible!
If you were adopted and don't have any known DNA relatives or genealogy information, then you will need to establish possible relationships with matches. A match with Gen = 4 means you probably share the same great-great-grandfather. Collect surname lists and family trees for matches. Continue to connect with additional cousins to find common surnames. Triangulate with multiple cousins to add credibility to your possible relationships - if two of your matches both share the same great-great-grandfather, that person or someone on his line is probably your ancestor. As you weave together possible relationships, you may discover intersections such as two different possible great-great-grandfathers who had grandchildren who married each other - they are probably your grandparents!
This is essential genealogy research in reverse - instead of trying to expand your lines and find new distant ancestors, you want to discover multiple possible ancestors and try to find where they or their descendants intersect.